Maximizing Production Efficiency in a Chemical Plant¶
In this section, we implement a black-box optimization example — originally introduced in the Amplify SDK demos and tutorials — using Amplify-BBOpt.
The sample code we consider here is shown below. For detailed explanations of the simulator and problem setup, please refer to the following demo tutorial page:
Black-Box Optimization of Operating Condition in a Chemical Reactor In this tutorial, the goal is to optimize the operating condition of a plant reactor to maximize production. The chemical reaction and transport phenomena of reactants and products in the reactor are predicted by numerical simulations using the finite difference method. Based on the predicted results, optimization is performed to maximize the amount of material production in the reactor.
Implementation of the Reactor Simulation¶
Below is the implementation of the Reactor simulation class based on the finite difference method (FDM). This implementation is essentially identical to that used in the demo mentioned above.
Implementing the black-box function¶
Here, we implement the black-box function using the reactor simulator class Reactor together with the Amplify-BBOpt @blackbox decorator.
from logging import getLogger
from typing import Annotated
from amplify_bbopt import AMPLIFY_BBOPT_LOGGER_NAME, BinaryVariable, blackbox
logger = getLogger(AMPLIFY_BBOPT_LOGGER_NAME)
input_q = [BinaryVariable() for _ in range(100)] # List of binary variables
# Objective function (returns the negative of the total amount of product B)
@blackbox
def my_obj_func(init_a: Annotated[list[int], input_q]) -> float:
my_reactor = Reactor()
minus_total = -my_reactor.integrate(init_a, fig=False)
logger.info(f"{minus_total=:.2e}, {my_reactor.cpu_time=:.1e}s")
return minus_total
Running the optimization¶
In this example, we use a Factorization Machine (FM) as the surrogate model via FMTrainer and instantiate the optimizer as FMQA. For other surrogate models, see the Model Trainer section of the documentation.
After adding the initial training data, we run the optimization cycles.
from datetime import timedelta
from amplify import FixstarsClient
from amplify_bbopt import FMTrainer, Optimizer
client = FixstarsClient()
client.parameters.timeout = timedelta(milliseconds=2000)
# client.token = "xxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxx" # Enter your Amplify AE access token when using a local environment
optimizer = Optimizer(my_obj_func, FMTrainer(), client)
num_init_data = 10 # Number of initial training samples
optimizer.add_random_training_data(num_init_data)
optimizer.optimize(10)
Plotting the Optimization History¶
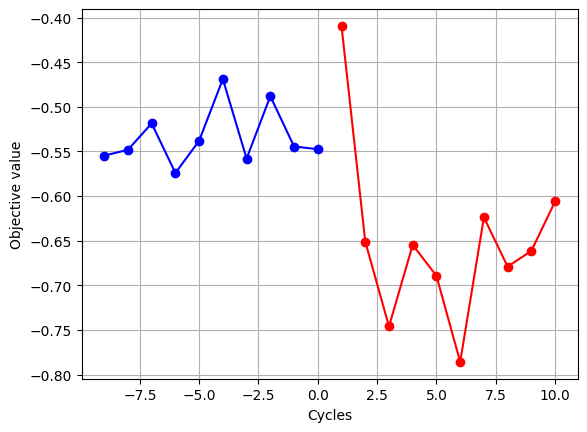
Due to the heuristic nature of the algorithms used by many Ising machines, the obtained solutions are not perfectly reproducible. Below, we show a typical example of the standard output and figure generated when running this sample code.
We were able to obtain optimization results consistent with those shown in the corresponding tutorial code.
# Objective values from the initial training data
objective_init = optimizer.training_data.y[:num_init_data]
# Objective values of the best solutions directly obtained from annealing
objective_anneal = [
float(h.annealing_best_solution.objective) for h in optimizer.history
]
plt.plot(range(-num_init_data + 1, 1), objective_init, "-ob")
plt.plot(range(1, len(objective_anneal) + 1), objective_anneal, "-or")
plt.xlabel("Cycles")
plt.ylabel("Objective value")
plt.grid(True)
Hint
In the current black-box function, the range of function values is clearly \(-1 \le y\le0\). To improve optimization performance, when applying the transformation described in this section, it is recommended to apply an offset before the exponential transformation so that the target objective value becomes zero.